Review Article - Onkologia i Radioterapia ( 2023) Volume 17, Issue 2
Artificial intelligence, machine learning and deep learning in neuroradiology: Current applications
Danilo Caudo*, Alessandro Santalco, Simona Cammaroto, Carmelo Anfuso, Alessia Biondo, Rosaria Torre, Caterina Benedetto, Annalisa Militi, Chiara Smorto, Fabio Italiano, Ugo Barbaro and Rosa MorabitoDanilo Caudo, IRCCS Centro Neurolesi Bonino-Pulejo (Messina - ITALY), Italy, Email: danycaudo@gmail.com
Received: 04-Jan-2023, Manuscript No. OAR-23-86840; Accepted: 25-Jan-2023, Pre QC No. OAR-23-86840 (PQ); Editor assigned: 06-Jan-2023, Pre QC No. OAR-23-86840 (PQ); Reviewed: 20-Jan-2023, QC No. OAR-23-86840 (Q); Revised: 22-Jan-2023, Manuscript No. OAR-23-86840 (R); Published: 01-Feb-2023
Abstract
Artificial intelligence is rapidly expanding in all medical fields and especially in neuroimaging/neuroradiology (more than 5000 articles indexed on PubMed in 2021) however, few reviews summarize its clinical application in diagnosis and clinical management in patients with neurological diseases. Globally, neurological and mental disorders impact 1 in 3 people over their lifetime, so this technology could have a strong clinical impact on daily medical work. This review summarizes and describes the technical background of artificial intelligence and the main tools dedicated to neuroimaging and neuroradiology explaining its utility to improve neurological disease diagnosis and clinical management.
Keywords
deep learning, artificial intelligence, machine learning, neuroradiology, neuroimaging
Introduction
The ever-increasing number of diagnostic tests requires rapid reporting without reducing diagnostic accuracy [1], and this could lead to misdiagnosis. In this context, the recent exponential increase in publications related to Artificial Intelligence (AI) and the central focus on artificial intelligence at recent professional and scientific radiology meetings underscores the importance of artificial intelligence to improve neurological disease diagnosis and clinical management.
Currently, there are many well-known applications of AI in diagnostic imaging, however, few reviews summarized its applications in Neuroimaging/neuroradiology. Thus, we aim at providing a technical background of AI and an overview of the current literature on the clinical applications of AI in neuroradiology/neuroimaging highlighting current tools and rendering a few predictions regarding the near future.
Technical background of AI
Any computer technique that simulates human intelligence is considered AI. AI is composed of Machine Learning (ML) and Deep Learning (DL) (Figure 1). ML designs systems to learn and improve from experience without being preprogrammed based on statistical data using computer technology. ML uses observations and data which are taken as examples to create some models and algorithms which are then used to make future decisions. In ML, some “ground truth” exists, which is used to train the algorithms. One example is a collection of brain CT scans that a neuroradiologist has classified into different groups (ie, haemorrhage versus no haemorrhage). The goal is to design software to learn automatically and independently of any human intervention or assistance for further intended decisions (Figure 2). DL, representing ML processing, instead applies artificial Convolutional Neural Networks (CNNs) to accelerate the learning process [2, 3]. CNNs are non-linear structures of statistical data organized as modelling tools. They can be used to simulate complex relationships between inputs and outputs using several steps (layers) of nonlinear transformations, which other analytic functions cannot represent [2].
Figure 1: AI uses computers to mimic human intelligence
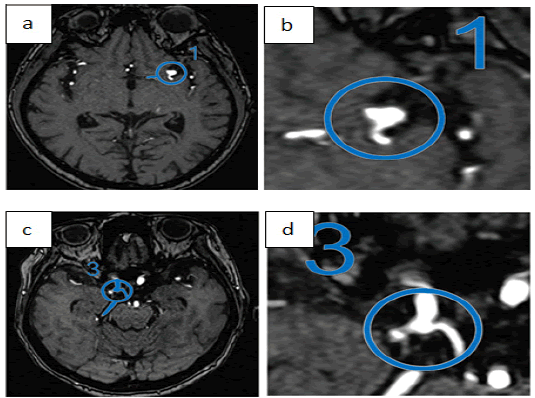
Figure 2: Automated Aneurysm detection on time-of-flight MRI.Ueda et al. [44]
CNNs can be trained to classify an image based on its characteristics through the observation of different images. More specifically, DL can identify common features in different images to use them as a classification model. For example, DL can be trained to find common features in variable images with and without a given pathology to discriminate between both entities. Consequently, it is possible to determine a specific diagnosis without human intervention and therefore there is some potential to improve both the time efficiency and th e productivity of ra diologists. A strength of DL is that its learning is based on growing experience. As a result, DL has enormous potential because it could update its response models by collecting data from large databases such as the Internet or the Picture Achieving and Comunication System (PACS). However, a limit is that as a consequence, algorithm performance depends largely on both the quantity and quality of data on which it is trained [4]. For example, an algorithm for tumour detection trained on a data set in which there is no occipital tumour is likely to have a higher error rate for tumours in that location. For a more complete description of DL, the reader is directed to the paper by Montagnon et al. [5].
Image acquisition and image quality improvement
Deep learning methods can be used to perform image reconstruction and improve image quality. AI can "learn" standard MR imaging reconstruction techniques, such as Cartesian and non-Cartesian acquisition schemes [6]. Additionally, deep learning methods could be applied to improve image quality. If low and high-resolution images are available, a deep mesh can be used to improve the resolution [7]. This has already been applied to CT imaging to improve resolution in low-dose CT images [8]. Another approach to improve image quality is to use MR images acquired at different magnetic fi eld strengths and co upled from the same anatomy [9].
AI is also able to reduce image acquisition times, this is especially useful in the case of DTI sequences, where the need for more angular directions extends the examination beyond what many patients can tolerate. A deep learning approach can reduce imaging duration by 12 times by predicting final parameter maps (fractional anisotropy, mean diffusivity, and so on) from relatively few angular directions [10]. There a re a lso s tudies i n which DL has increased the signal-to-noise ratio in Arterial Spin-Labelling (ASL) sequences to improve image quality [11]. Finally, some applications of AI could improve resolution and image enhancement by providing a better resolution and signal-to-noise ratio reducing the dose of contrast needed to provide diagnostic images [2].
Clinical AI applications in neuroradiology/ neuroimaging
Recently, 37 AI applications were reviewed in the domain of Neuroradiology/Neuroimaging from 27 vendors offering 111 functionalities [12]. These AI functionalities mostly support radiologists and extend their tasks. Interestingly, these AI applications are designed for just one pathology, such as ischemic stroke (35%), intracranial haemorrhage (27%), dementia (19%), multiple sclerosis (11%), or brain tumour (11%) to mention the most common [12]. In our review, we found miscellaneous clinical applications of AI in neuroradiology/neuroimaging ranging from the detection and classification of anomalies on imaging to the prediction of outcomes with disease quantification by estimating the volume of anatomical structures, the burden of lesions, and the volume of the tumour. In particular, regarding detecting tools, primary emphasis has been placed on identifying urgent findings that enable worklist prioritization for abnormalities such as intracranial haemorrhage [13-19], acute infarction [20-23], large-vessel occlusion [24, 25], aneurysm detection [26-28], and traumatic brain injury [29-31] on non-contrast head CT.
Other AI detecting tools are in brain degenerative disease, epilepsy, oncology, degenerative spine disease (to detect the size of the spinal canal, facet joints alterations, disc herniations, size of conjugation foramina, and in scoliosis the Cobb angle), fracture detection (vertebral fracture such as compression fracture), and in multiple sclerosis to identify disease burden over time and predicting disease activity. In glioma, some DL algorithms were tested to predict glioma genomics [32-36]
Regarding segmentation tools, we found tools able to segment vertebral disc, vertebral neuroforamina, and vertebral body for degenerative spine disease, brain tumour volume in the neurooncological field, and white and grey matter in degenerative brain diseases.
In the following paragraphs, we report the main diseases where AI is useful with the more significant relative studies.
Intracranial haemorrhage:
Intracranial haemorrhage detection has been widely studied as a potential clinical application of AI [37-39], able to work as an early warning system, raise diagnostic confidence [40], and classify haemorrhage types [41]. In particular, in these studies, Kuo et al. developed a robust haemorrhage detection model with an area under the receiver operating characteristic curve (ROCAUC) of 0.991 [14]. They trained a CNN on 4396 CT scans for classification and segmentation concurrently on training data which were labelled pixel-wise by attending radiologists. Their test set consisted of 200 CT scans obtained at the same institution at a later time. They report a sensitivity of 96% and a specificity of 98%.
Ker et al. developed a 3-Dimensional (3D) Convolutional Neural Network (CNN) model to detect 4 types of intracranial haemorrhage (subarachnoid haemorrhage, acute subdural haemorrhage, intraparenchymal hematoma, and brain polytrauma haemorrhage) [41]. They used a data set consisting of 399 locally acquired CT scans and experimented with data augmentation methods as well as various threshold levels (i.e, window levels) to achieve good results. They measured performance as a binary comparison between normal and one of the 4 haemorrhage types and achieved ROC-AUC values of 0.919 to 0.952. The RSNA 2019 Brain CT Hemorrhage Challenge was another milestone, in which a data set of 25 312 brain CT scans were expert-annotated and made available to the public [42]. The scans were sourced from 3 institutions with different scanner hardware and acquisition protocols. The submitted models were evaluated using logarithmic loss, and top models achieved excellent results on this metric (0.04383 for the first-place model). However, it is difficult to di rectly compare th is re sult to other st udies th at utilize the ROC-AUC of haemorrhage versus no haemorrhage as their performance metric. Approved commercial software for haemorrhage detection now exist on the market and has been evaluated in clinical settings. Rao et al. evaluated Aidoc (version 1.3, Tel Aviv) as a double reading tool for the prospective review of radiology reports [40]. They assessed 5585 non-contrast CT scans of the head at their institutions which were reported as being negative for haemorrhage and found 16 missed haemorrhages (0.2%), all of which were small haemorrhages. The software also flagged 12 false positives. The Aidoc software was also tested as a triage tool by Ginat in which the software, evaluating 2011 non-contrast CT head scans, contained both false positive and false negative findings of haemorrhage [13]. The st udy re ports sensitivity and specificity of 88.7% and 94.2% for haemorrhage detection, respectively. The author however described a benefit of false-positive flags for haemorrhage as these studies sometimes contained other hyperattenuating pathologies. On the flip side, the author reports a drawback in flagging in patient scans in which a haemorrhage is stable or even improving, which may unnecessarily prioritize nonurgent findings.
Aneurysm detection:
Detecting unruptured intracranial aneurysms has significant clinical importance considering that they account approximately for 85% of non-traumatic subarachnoid haemorrhages and their prevalence is estimated at approximately 3% [43]. MRI with Time- Of-Flight Angiography sequences (TOF-MRA) is the modality of choice for aneurysm screening, as it does not involve ionizing radiation nor intravenous contrast agents. Deep learning has been used to detect aneurysms on TOF-MRA. Published methods demonstrate high sensitivity but poor specificity, resulting in multiple false positives per case. Although this fact necessitates a close review of all aneurysms flagged by the software, the resulting models may nevertheless be useful as screening tools. Ueda et al. [44], tested a DL model on 521 scans from the same institution as well as 67 scans from an external data set. DL model achieved 91% to 93% sensitivity with a rate of 7 false positives per scan. Despite the high false positive rate, the authors found that this model helped them detect an additional 4.8% to 13% of aneurysms in the internal and external test data sets (Figure 2). In another study, Faron et al., trained and evaluated a CNN model on a data set of TOF-MRA scans on 85 patients. CNN achieved 90% sensitivity with a rate of 6.1 false positives per case [45]. Similarly, Nakao et al. trained a CNN on a data set of 450 patients with TOF-MRA scans on 3-Tesla magnets only [46]. They were able to achieve a better result at 94% sensitivity with 2.9 false positives per case. Yang et al. used a different approach, in which they produced 3D reconstructions of the intracranial vascular tree using TOF-MRA images [47]. They annotated aneurysms on the 3D projections and used them to train several models. In this manner, they achieved a good discriminatory result between healthy vessels and aneurysms.
Stroke:
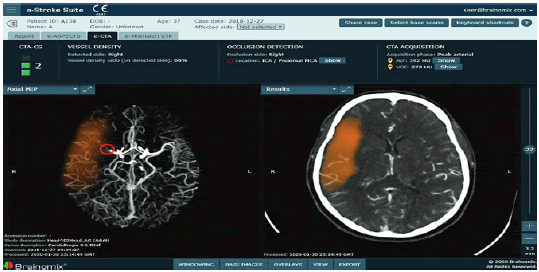
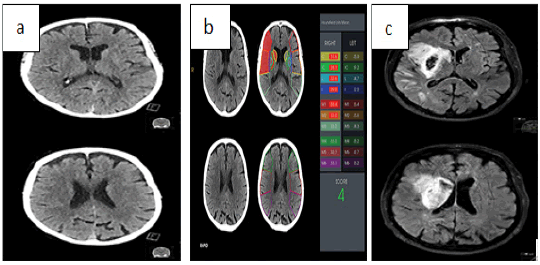
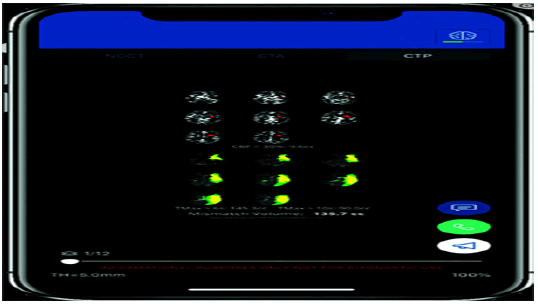
In stroke imaging 3 components of stroke imaging are explored: Large Vessel Occlusion (LVO) detection (Figure 3), automated measurement of core infarct volume and the Alberta Stroke Program Early CT Score (ASPECTS) (Figure 4), and infarct prognostication (Figure 5).
Figure 3: Brainomix e-CTA tool demonstrating identification and localization of an LVO of the right MCA, collateral score and collateral vessel attenuation, and a heat map of the collateral deficit (orange)
Figure 4: A 74-year-old with a right-sided stroke due to MCA occlusion a) Two neuroradiologists assessed ASPECTS of 9 on the initial CT images b) Rapid aspects software evaluation is 4 on the same initial CT images
Figure 5: AI mobile interface showing a left MCA territory infarction with a mismatch on perfusion CT [59]
The timely detection of STROKE is critical in brain ischemic treatment, in this context AI has shown the potential in reducing the time to diagnosis. In particular, there are some AI applications able to detect LVO. You et al. developed an LVO detection model using clinical and imaging data (non-contrast CT scans of the head) [48]. AI detects the hyperdense Middle Cerebral Artery (MCA) sign which is a finding suggestive of the presence of an MCA thrombus. The AI, based on the U-Net architecture, was trained and tested on a local data set of 300 patients. It achieved a sensitivity and specificity of 68.4% and 61.4%, respectively. On NCCT, an SVM algorithm detected the MCA dot sign in patients with acute stroke with high sensitivity (97.5%) [49]. A neural network that incorporated various demographic, imaging, and clinical variables in predicting LVO outperformed or equalled most other prehospital prediction scales with an accuracy of 0.820 [50]. A CNN-based commercial software, Viz-AI-Algorithm v3.04, detected proximal LVO with an accuracy of 86%, a sensitivity of 90.1%, a specificity of 82.5, AUC of 86.3% (95% CI, 0.83-0.90; P # .001), and intraclass correlation coefficient (ICC) of 84.1% (95% CI, 0.81-0.86; P # .001), and Viz-AI-Algorithm v4.1.2 was able to detect LVO with high sensitivity and specificity (82% and 94%, respectively) [51]. Unfortunately, no study has yet shown whether AI methods can accurately identify other potentially treatable lesions such as M2, intracranial ICA, and posterior circulation occlusions.
Establishing infarct volumes is important to triage patients for appropriate therapy. AI has been able to establish core infarct volumes on MRI sequences through automatic lesion segmentation [52-54]. One reported limitation was the reliance on FLAIR and T1 images that do not fully account for the timing of stroke occurrence. Another limitation was a tendency to overestimate the volume of small infarcts and underestimate large infarcts compared with manual segmentation by expert radiologists and difficulty in distinguishing old versus new strokes [54]. Discrepancies in volumes were attributed to nondetectable early ischemic findings, partial volume averaging, and stroke mimics on CT [55].
ASPECTS is an important early predictor of infarct core for middle cerebral artery (MCA) territory ischemic strokes [56]. It assesses 10 regions within the MCA territory for early signs of ischemia and the resulting score ranges from 0 to 10, where 10 indicates no early signs of ischemia, while 0 indicates ischemic involvement in all 10 regions. The score is currently a key component in the evaluation of the appropriateness of offering endovascular thrombectomy. Several commercial AI applications perform automated ASPECTS evaluation and they have been assessed in clinical settings. In particular, Goebel et al. compared Frontier ASPECTS Prototype (Siemens Healthcare GmbH) and e-ASPECTS (Brainomix) to 2 experienced radiologists and found that e-ASPECTS showed a better correlation with expert consensus [57]. Guberina et al. compared 3 neuroradiologists with e-ASPECTS and found that the neuroradiologists had a better correlation with infarct core as judged on subsequent imaging than the software [58]. Maegerlein et al. compared RAPID ASPECTS (iSchemaView) to 2 neuroradiologists and found that the software showed a higher correlation with expert consensus than each neuroradiologist individually [59]. An example output of RAPID ASPECTS is shown in Figure 2. Accuracy varies widely and depends on the software and chosen ground truth. An interesting result suggested by one study, however, was that the RAPID software produced more consistent results when the image reconstruction algorithm was varied compared to human readers [60]. The i nterclass c orrelation c oefficient bet ween multiple reconstruction algorithms was 0.92 for RAPID, 0.81 to 0.84 for consultant radiologists, and 0.73 to 0.79 for radiology residents.
Prognostication in Stroke treatment is critical to detect patients who are most likely to benefit f rom t reatment c onsidering t he risks related. For this reason, AI has been studied as a tool for predicting post-treatment outcomes. In this context developed a CNN model to predict post-treatment infarct core based on initial pre-treatment Magnetic Resonance Imaging (MRI) [23]. The authors used a locally acquired data set of 222 patients, 187 of whom were treated with tPA. The model was evaluated using a modified version of the ROC-AUC, where the true positive rate was set to the number of voxels correctly identified as positive, the true negative rate was set to the number of voxels correctly identified as negative, and so on. The reported modified ROC-AUC is 0.88. In another study, Nishi et al. developed a U-Net model to predict clinical post-treatment outcomes using pretreatment diffusion-weighted imaging on patients who underwent mechanical thrombectomy [61]. The clinical outcome was defined using the modified Rankin Sc ale (mRS) at 90 days after the stroke. The outcomes were categorized as ‘‘good’’ (mRS < 2) and ‘‘poor’’ (mRS > 2). After training on a data set of 250 patients, the model was validated on a data set of 74 patients and found to have a ROC-AUC of 0.81.
Multiple sclerosis:
In multiple sclerosis deep learning has been investigated as a tool to estimate disease burden and predict disease activity through MRI imaging. MRI is used to assess disease burden over time but this requires comparison with prior scans, which can be burdensome and error-prone when the number of lesions is large. Nair et al. evaluated a DL algorithm for MS lesion identification on a private multicenter data set of 1064 patients diagnosed with the relapsingremitting variant, containing a total of 2684 MRI scans. In this study, the DL performance was worst with small lesions [62]. The algorithm was tested on 10% of the data set where it achieved a ROC-AUC of 0.80 on lesion detection. In another study, Wang et al. trained a CNN on 64 Magnetic Resonance (MR) scans to detect MS lesions which were able to achieve a sensitivity of 98.77% and specificity of 98.76% for lesion detection, respectively [63].
Regarding predicting disease activity, Yoo et al. developed a CNN that combined a data set of 140 patients, who had onset of the first demyelinating symptoms within 180 days of their MR scan, with defined clinical measurements [64]. CNN achieved a promising result with a ROC-AUC of 0.746 for predicting progression to clinically definite MS.
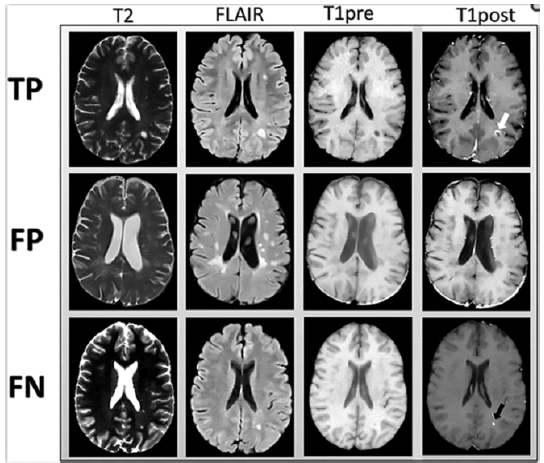
Another application of AI in MS regards efforts made to reduce gadolinium use where possible considering emerging evidence that repeated administrations of gadolinium-based contrast agents lead to their deposition in the brain [65]. In particular, Narayana et al. [66] used DL to predict lesion enhancement based on their appearance on non-contrast sequences (precontrast T1-weighted imaging, T2-weighted imaging, and fluid-attenuated inversion recovery) (Figure 6). They used a data set of 1008 patients with 1970 MR scans acquired on magnets from 3 vendors. DL achieved a ROC-AUC of 0.82 on lesion enhancement prediction, suggesting that this approach may help reduce contrast use.
Figure 6: Examples of images input to the network (T2-weighted, fluidattenuated inversion recovery and pre-contrast T1-weighted images). Post-contrast T1-weighted images demonstrating areas of true-positive (white arrow) and false-negative (black arrow) enhancement are shown for comparison [66]
Fracture detection:
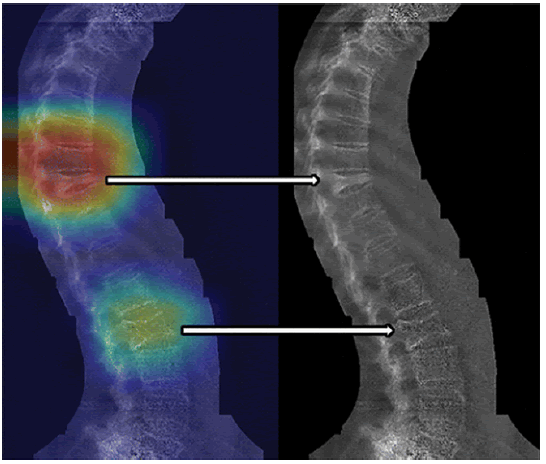
Regarding fracture detection tools we found that Tomita et al. tested a DL model to detect osteoporotic vertebral fractures in a data set of 1432 CT scans. The outcome was a binary classification of whether or not a fracture was present [67]. Using 80% of the data set for training, a ROC-AUC of 0.909 to 0.918 was achieved with an accuracy of 89.2%. This was found to be equivalent to radiologists on the same data set. In a similar study by Bar et al. a CNN was trained with a data set of 3701 CT scans of the chest and/or abdomen to detect vertebral compression fractures. The model was able to detect vertebral compression fractures with 89.1% accuracy, 83.9% sensitivity, and 93.8% specificity [68]. Furthermore, [69] used a data set of 12 742 dual-energy X-ray absorptiometry scans to train a binary classifier for the detection of vertebral compression fractures (Figure 7); 70% of the data set was used for training, which yielded a ROC-AUC of 0.94. The optimal threshold achieved a sensitivity of 87.4% and a specificity of 88.4%.
Figure 7: Images of a 77-year-old female patient evaluated for vertebral fracture. Hear map shows one severe vertebral compression fracture (upper arrow) and one mild fracture (lower arrow). Heat maps are unitless low-resolution images showing relative contributions of general areas in images to the prediction. The heat map has been overlaid on the original vertebral fracture assessment image (left side). Arrows denote the corresponding locations of vertebral fractures on the original images as presented to the convolutional neural network (right side)[69]
Brain tumour:
For brain tumours there AI application for segmentation, that can be used as a stand-alone clinical tool, such as in contouring targets for radiotherapy, or it can also be used to extract tumours as a preliminary step for further downstream ML tasks, such as diagnosis, pre-surgical planning, follow-up and tumour grading [16, 70-74]. Unfortunately, there is a limit to the AI segmentation, usually, only a minority of voxels represent tumours and the majority represent healthy tissue, however, in a recently published study, Zhou et al. trained an AI model with a publicly available MRI data set of 542 glioma patients and they were able to tackle this limit [71]. Their results demonstrate excellent performance with a Dice score of 0.90 for the whole tumour (entire tumour and white matter involvement) and 0.79 for tumour enhancement.
Another AI clinical application for brain tumours is predicting glioma genomics Isocitrate Dehydrogenase (IDH) mutations that are important prognosticators [75, 76]. In 2019 multiple studies investigated the prediction of IDH mutation status from MRI. Zhao et al. published a meta-analysis of 9 studies totalling 996 patients . The largest data set used for training had 225 patients. These studies developed binary classification models and ha d a ROC-AUC of 0.89 (95% CI: 0.86-0.92). Pooled sensitivities and specificities were 87% (95% CI:76-93) and 90% (95% CI:72-97), respectively. Since then, another study was published by Choi et al. [34] using a larger MRI data set of 463 patients. It showed excellent results with ROC-AUC, sensitivity, and specificity of 0.95, 92.1%, and 91.5%, respectively. This model used a CNN to segment and as a feature extractor to predict IDH mutation risk. Haubold et al. used 18F-fluoro-ethyl-tyrosine Positron Emission Tomography (PET) combined with MRI to predict multiple tumour genetic markers using a data set of 42 patients before biopsy or treatment [77]. They trained 2 different classical ML models and used biopsy results as the ground truth. They achieved a ROC-AUC of 0.851 for predicting the ATRX mutation, 0.757 for the MGMT mutation, 0.887 for the IDH1 mutation, and 0.978 for the 1p19q mutation.
Degenerative brain disease:
Regarding dementias numerous AI networks have been trained from large longitudinal datasets such as the Alzheimer’s Disease Neuroimaging Initiative (ADNI), resulting in many diagnostic DL tools for Alzheimer's Disease (AD), such as models using 18F Fluoro-Deoxy-Glucose (FDG) PET76 and structural MRI of the hippocampus to predict AD onset from 1 years to 6 years in advance [78]. Furthermore, AI may assist in the diagnosis of dementia types. AI can differentiate AD from Lewy body and Parkinson’s dementia [32]. Similarly, other AI tools can differentiate between Mild Cognitive Impairment (MCI) and AD [79]. In addition to diagnosis, AI can also probe neurobiology. New ML techniques such as Subtype and Stage Inference have provided novel neuroimaging and genotype data-driven classifications of diagnostic subtypes and progressive stages for AD and Fronto Temporal Dementia (FTD). SuStaIn has localized distinct regional hotspots for atrophy in different forms of familial FTD caused by mutations in genes [80].
In other studies, DL has integrated MRI, neurocognitive, and APOE genotype information to predict conversion from MCI to AD [81]. Combining several AI systems (including structural MRI and amyloid PET) may augment the diagnosis and management of the complex natural history of AD. In the future, the integration of AI tools for imaging with AI systems designed to examine serum amyloid markers mortality prediction from clinicians’ progress notes and assessments of cognition, and postmortem immunohistochemistry images [82-85], may improve many facets of care in neurodegenerative disease. In Huntington's Disease, an autosomal dominant movement disorder, diagnosis, and management may be enhanced by incorporating CAG repeat length data with CNN developed for caudate volumetry, and objective gait assessment [86]. Such multi-approaches may improve risk stratification, p rogression monitoring, a nd c linical management in patients and families.
Epilepsy:
The u se o f A I i n the d iagnosis o f e pilepsy c ould i mprove the diagnostic capabilities of this condition as the symptoms are not specific and often overlap with other conditions [87, 88].. In particular, the integration of anamnestic, clinical, electroencephalographic and imaging information is fundamental for an accurate diagnosis and subtype differentiation [89]. Neuroimaging plays an important role in both diagnosis and follow-up and prognosis. In particular, structural Magnetic Resonance Imaging (sMRI) can help identify cortical abnormalities (e.g. temporal mesial sclerosis, Focal Cortical Dysplasia [FCD], neoplasms, etc.), while functional Magnetic Resonance Imaging (fMRI), emission tomography positron imaging and Magneto-Encephalo-Graphy (MEG) can help localize brain dysfunction.
Park et al. (2020) used an SVM classifier on bilateral hippocampi [90]. The model obtained an area under the receiver operating characteristic curve (AUC) of 0.85 and an accuracy of 85% in differentiating e pileptic p atients f rom h ealthy c ontrols, b etter than human evaluators. Mesial sclerosis is often subtle and invisibile. Such cases can lead to a misdiagnosis and consequently delay the surgical treatment. Therefore, recent machine learning models have been proposed to identify MRI-negative patients and lateralize foci. For example, Mo et al. (2019) used an SVM classifier based o n clinically e mpirical f eatures, a chieving 8 8% accuracy in detecting MRI-negative patients and an AUC of 0.96 in differentiating MRI-negative patients from controls [91]. The most important feature was the degree of blurring of the greywhite matter at the temporal pole. Similarly, Beheshti et al. (2020) used an SVM to diagnose epileptic patients for mesial sclerosis and lateralize foci in a cohort of 42 MRI-negative PET-positive patients [92]. Focusing on FLAIR, a simple and widely available sequence, the authors extracted signal strength from Regions Of Interest (ROIs) a priori. The model achieved 75% accuracy in differentiating right and left epileptics from controls. The best performance was obtained in identifying right epilepsy, with an accuracy of 88% and an AUC of 0.84. The most important ROIs were the amygdala, the inferior, middle and superior temporal gyrus and the temporal pole.
However, analyzing only the temporal lobes may not reveal a more global pathology, for this reason, Sahebzamani et al. (2019), using unified segmentation and an SVM classifier, found that whole brain features are more diagnostic than hippocampal features alone (94% vs 82% accuracy) [93]. In particular, global contrast and white matter homogeneity were found to be the most important, along with the clustering tendency and grey matter dissimilarity. In particular, the best performances were obtained based on the mean sum of the whole brain’s white matter.
Another possibility of AI on sMRI is to lateralize the temporal epileptic focus. In a study with an SVM, the combination of hippocampus, amygdala, and thalamic volumes was more predictive of the laterality of the epileptic focus. The combined model achieved 100% accuracy in patients with mesial sclerosis (Mahmoudi et al., 2018) [94]. Furthermore, Gleichgerrcht et al. (2021) used SVM deep learning models to diagnose and lateralize temporal epileptic focus based on structural and diffusionweighted MRI ROI data [95]. The models achieved an accuracy of 68%-75% in diagnosis and 56%-73% in lateralization with diffusion data. Based on the sMRI data, ipsilateral hippocampal volumes were the most important for prediction performance. Based on the dwMRI data, ipsilateral tracked beams had the highest predictive weight.
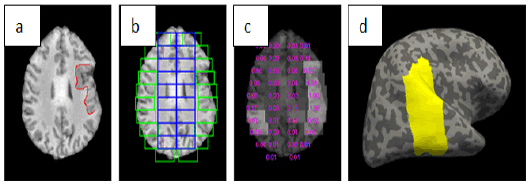
Machine learning techniques have also been used to diagnose cortical dysplasia, the most common cause of medically refractory epilepsy in children and adults second most common cause (Kabat and Król, 2012) [96]. In one study, Wang et al. (2020) trained a CNN to exploit the differences in texture and symmetry on the boundary between white matter and grey matter (Figure 8) [97]. The model achieved a diagnostic accuracy of 88%. Similarly, Jin et al. (2018) trained surface-based morphometry and a non-linear neural network model [98]. Based on six characteristics of a 3D cortical reconstruction, the model achieved an AUC of 0.75. The contrast of intensity between grey matter and white matter, local cortical deformation and local cortical deformation of cortical thickness were the most important factors for classification. Notably, the model worked well in three independent epilepsy centres. However, data extraction limits have been reported regarding variations in image quality between different clinical sites of the tests.
Figure 8: An example of FCD detection a) an axial slice with FCD lesion labelled in red b) patch extraction results c) classification results and numbers stand for probabilities of being FCD patches d) detection mapped onto the inflated cortical surface (shown in yellow) [97]
Another possibility is to combine features derived from sMRI with those derived from fMRI for a variety of clinical applications, such as diagnosing epilepsy and predicting the development of epilepsy following traumatic events. In one study, Zhou et al. (2020) found that the combination of fMRI and sMRI functions was more useful than either modality alone in identifying epileptic patients [99]. In another study, Rocca et al. (2019) used random forest and SVM models to help predict the development of seizures following temporal brain injuries [100]. The highest AUC of 0.73 was achieved with a random forest model using functional characteristics. Therefore, additional studies that directly compare the additive utility of fMRI and sMRI, perhaps using framework models, may be useful.
In summary, a variety of machine learning approaches have been used for the automated analysis of sMRI data in epilepsy. Given the limited number of publicly available sMRI datasets, models tend to be trained on small single-centre cohorts, this and the lack of external validation limits the interpretation of widespread clinical utility. Future work with larger, multicenter data sets is needed.
Functional MRI, a method that measures changes in blood flow to assess and map the magnitude and temporospatial characteristics of neural activity, is gaining popularity in the field of epilepsy [101]. A growing body of evidence suggests that epilepsy is likely characterized by complex and dynamic changes in the way neurons communicate (i.e., changes in neural networks and functional connectivity), both locally and globally. AI applied to fMRI is useful in recent epilepsy studies. Mazrooyisebdani et al. (2020) used an SVM to diagnose temporal epilepsies based on functional connectivity characteristics derived from graph theory analysis [102]. The model achieved an accuracy of 81%. Similarly, Fallahi et al. (2020) constructed static and dynamic matrices from fMRI data to derive measurements of global graphs [33]. Then, the most important characteristics were selected using random forest and the classification was performed with SVM. The use of dynamic features led to better accuracy than the use of static features (92% versus 88%) in the lateralization of temporal epilepsy. In another study, Hekmati et al. (2020) used fMRI data to quantify mutual information between different cortical regions and insert these quantities into a four-layered perceptual classifier [103]. The model achieved 89% accuracy in locating seizure foci. Finally, Hwang et al. (2019) used LASSO feature selection to extract functional connectivity features, which were then used to train an SVM, linear discriminant analysis, and naive Bayes classifier to diagnose temporal epilepsy [104]. The best accuracy of 85% was achieved with the SVM model. Recent work by Bharat et al. (2019) with machine learning provided further evidence that epilepsy arises from impaired functional neural networks. Using resting-state fMRI (rs-fMRI), the researchers were able to identify connectivity networks specific to temporal epilepsy [105]. The model differentiated temporal epilepsy patients from healthy controls with 98% accuracy and 100% sensitivity. The networks were also found to be highly correlated with diseasespecific clinical features and hippocampal atrophy. Although this evidence provides new proof of concept for the existence of specific epilepsy networks, future work is needed, as impaired functional activity can occur secondary to the effects of antiepileptic drugs or a variety of other confounding factors.
In summary, fMRI-based machine learning can be used to identify complex alterations in functional neural networks in the epileptic brain and further exploit these differences for classification purposes. In many cases of epilepsy, structural and functional anomalies of the network probably coexist. Current machine learning models with fMRI are limited by small sample sizes, probably because there are few publicly available data sets. However, fMRI is increasingly being integrated into routine clinical practice, particularly for lateralization before surgery [106]. Recent studies have shown that fMRI may be specifically useful in pre-surgical lateralization [107]. With technological advances and further methodological refinements, fMRI could become the standard of care in epilepsy and AI will be increasingly used to assist in diagnostic and prognostic tasks.
Another field of AI application is Diffusion Tensor Imaging (DTI), which has advantages in detecting subtle structural abnormalities of epileptogenic foci. Machine learning in DTI can use for classification improving the diagnosis and treatment of epilepsy, particularly when used for pre-surgical planning and postsurgical outcome prediction [108].
Degenerative Spine Disease:
The rate of MRI examinations is stressfully increased due to the significant number of patients suffering from degenerative spine disease [109]. Consequently, radiologists face a work overload who need to evaluate numerous parameters (size of the spinal canal, facet joints, disc herniations, size of conjugation foramina, etc) in all spinal levels in a short time. Accordingly, different DL and ML algorithms that can automatically classify spinal pathology may help to reduce patient waiting lists and examination costs. In this context, Jamaludin et al. evaluated an automatic disc disease classification system that yielded an accuracy of 96% compared to radiologist assessment. Notably, the main sources of limitation were either poor scan quality or the presence of transitional lumbosacral anatomy [109]. Furthermore, Chen et al. evaluated a DL tool to measure Cobb angles in spine radiographs for patients with scoliosis. They used a data set of 581 patients and were able to achieve a correlation coefficient of r ¼ 903 to 0.945 between the DL-predicted angle and the ground truth. [110]
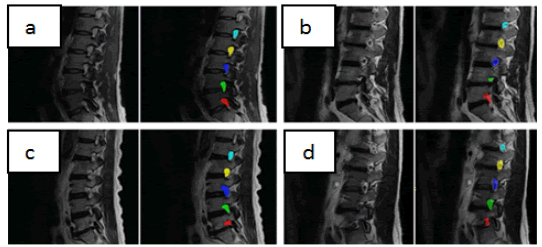
Regarding spine segmentation, some models have been developed with good results. Huang et al. achieved intersection over-union scores of 94.7% for vertebrae and 92.6% for disc segmentations on sagittal MR images using a training set of 50 subjects and a test set of 50 subjects [111]. Whitehead et al. trained a cascade of CNNs to segment spine MR scans using a data set of 42 patients for training and 20 patients for testing. They were able to achieve Dice scores of 0.832 for discs and 0.865 for vertebrae [112]. In this context DL has been used to answer research questions, for example, Gaonkar et al. used DL to look for potential correlations between the cross-sectional area of neural foramina and patient height and age, showing that the area of neural foramina is directly correlated with patient height and inversely correlated with age [35] (Figure 9).
Figure 9: A random sample of automated neural foraminal segmentations used for generating measurements. Original MRI Images (left) and overlaid computer-generated segmentation(right)
AI tools for ultrasound in neuroimaging
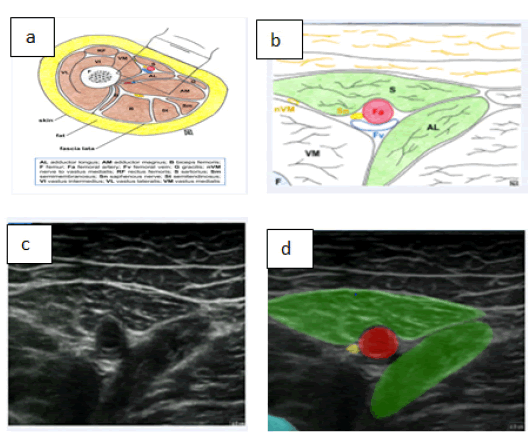
AI applications in neuroimaging and ultrasound (US) are mostly focused on the identification of anatomical structures such as nerves. A large number of algorithms have shown to be able to segment US images for these aims. For example, Kim et al. developed a neural network that accurately and effectively segments the median nerve. To train the algorithm and evaluate the model, 1,305 images of the median nerve of 123 normal subjects were used [113]. However, the proposed neural network yielded more accurate results in the wrist datasets, rather than forearm images, with a precision respectively of 90.3% and 87.8%. Different studies showed that AI may help to automatically segment nerve and blood vessels to facilitate ultrasound-guided regional anaesthesia [114–116]. Automated medical image analysis can be trained to recognize the wide variety of appearances of the anatomical structures and could be used to enhance the interpretation of anatomy by facilitating target identification (e.g., peripheral nerves and fascial planes) [117, 118]. For example, a model has been well developed for peripheral nerve block in the adductor canal. In this model, the sartorius and adductor longus muscles, as well as the femur, were first identified as landmarks. The optimal block site is chosen as the region where the medial borders of these two muscles align. The femoral artery is labelled as both a landmark and a safety structure. The saphenous nerve is labelled as a target. AI-applications assist the operator in identifying the nerve and the correct target site for the block (Figure 10) [118-120].
Figure 10: Sono-anatomy of the adductor canal block a) Illustration showing a cross-section of the mid-thigh b) Enlarged illustration of the structures seen on ultrasound during performance adductor canal block c) Ultrasound view during adductor canal block d) Ultrasound view labelled by AnatomyGuide [121]
Conclusion
This review explores important recent advances in ML and DL within neuroradiology. There have been many published studies exploring AI applications in neuroradiology, and the trend is accelerating. AI applications may cover multiple fields of neuroimaging/neuroradiology diagnostics, such as image quality improvement, image interpretation, classification of disease, and communication of salient findings to patients and clinicians. The DL tools show an outstanding ability to execute specific tasks at a level that is often compared to those of expert radiologists. In this context, AI may indeed have a role in enhancing radiologists’ performance through a symbiotic interaction which is going to be more likely mutualistic. However, the existing AI tools in neuroradiology/neuroimaging have been trained for single tasks so far. This means that an algorithm trained to detect stroke would not be able to show similar accuracy to detect and classify brain tumours and vice-versa. This is a great limit since patients often suffer from multiple pathologies, a complete AI assessment that integrates all different algorithms would be favourable. As far as we know, ML or DL models which are capable of simultaneously performing multiple interpretations have not yet been reported. We believe that this technology development may represent the key requirement to shift AI from an experimental tool to an indispensable application in clinical practice. AI algorithms for combined analysis of different pathologies should also warrant an optimized and efficient integration into the daily clinical routine. Furthermore, rigorous validation studies are still needed before these technological developments can take part in clinical practice, especially for imaging modalities such as MRI and CT, for which the accuracy of DL models highly depends on the type of scanner used and protocol performed. In addition, the reliability of AI techniques requires the highest validation also considering the legal liabilities that radiologists would hold for their usage and results.
Nowadays, only specific DL applications have demonstrated accurate performance and may be integrated into the clinical workflow under the supervision of an expert radiologist. In particular, AI algorithms for intracranial haemorrhage, stroke, and vertebral compression fracture identification may be considered suitable for application in daily clinical routines.
Other tasks, such as glioma genomics identification, stroke prognostication, epilepsy foci identification and predicting clinically definite MS, have shown significant progress in the research domain and may represent upcoming clinical applications in the not-so-distant future.
However, the majority of AI algorithms show that there is still a range of inaccuracies for example in labelling anatomical structures, especially in the context of atypical or complex anatomy. Moreover, another challenge will be to ensure the presence of highly skilled practitioners, since machine learning systems are not guaranteed to outperform human performance and these systems should not be relied upon to replace the knowledge of doctors.
Probably, the ongoing development of DL in neuroradiology/ neuroimaging will significantly influence the work of future radiologists and other specialists, which will need a specific AI education to begin during residential training, to deeply understand the mechanisms and potential pitfalls. Furthermore, knowledge of AI could be an opportunity to improve training in radiology and other specialities. AI can assign specific cases to trainees based on their training profile, to promote consistency in the trainees' individual experiences, and, in the context of anaesthetic procedures, to facilitate an easier understanding of anatomy.
Declarations
Funding
Current Research Funds 2023, Ministry of Health, Italy
Conflicts of interest/competing interests
The authors have no competing interests to declare relevant to this article’s content.
All authors certify that they have no affiliations with or involvement in any organization or entity with any financial interest or non-financial interest in the subject matter or materials discussed in this manuscript.
The authors have no financial or proprietary interests in any material discussed in this article.
Author Contributions
All authors whose names appear on the submission
1. Made substantial contributions to the conception and design of the work; and to the acquisition,analysis, and interpretation of data;
2. Drafted the work and revised it critically for important intellectual content;
3. Approved the version to be published; and
4. Agree to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
References
- McDonald RJ, Schwartz KM, Eckel LJ, Diehn FE, Hunt CH, et al.The effects of changes in utilization and technological advancements of cross-sectional imaging on radiologist workload. Acad radiol 2015;22:1191-1198.
- Hosny A, Parmar C, Quackenbush J, Schwartz LH, Aerts HJ. Artificial intelligence in radiology. Nat Rev Cancer. 2018;18:500-510.
- D’Angelo T, Caudo D, Blandino A. Artificial intelligence, machine learning and deep learning in musculoskeletal imaging: Current applications. J Clin Ultrasound JCU. 2022;50:1414-1431.
- Gebru T, Morgenstern J, Vecchione B, Vaughan JW, Wallach H. Datasheets for Datasets. Commun ACM. 2021;64:86-92
- Emmanuel M, Milena C, Alexandre CC, Hamilton V, Derennes T, et al. Deep learning workflow in radiology: a primer. Insights Imaging. 2020;11:22.
- Zhu B, Liu JZ, Cauley SF, Rosen BR, Rosen MS. Image reconstruction by domain-transform manifold learning. Nature. 2018;555:487-492.
- Dong C, Loy CC, He K, Tang X. Learning a Deep Convolutional Network for Image Super-Resolution. In European conf comput vis. 2014;184-199.
- Chen H, Zhang Y, Kalra MK, Lin F, Chen Y. Low-Dose CT With a Residual Encoder-Decoder Convolutional Neural Network. IEEE Trans Med Imaging. 2017; 36:2524-2535.
- Bahrami K, Shi F, Zong X, Shin HW, An H. Reconstruction of 7T-Like Images From 3T MRI. IEEE Trans Med Imaging. 2016;35:2085-2097.
- Golkov V, Dosovitskiy A, Sperl JI, Menzel MI, Czisch M. q-Space Deep Learning: Twelve-Fold Shorter and Model-Free Diffusion MRI Scans. IEEE Trans Med Imaging. 2016;35:1344-1351.
- Cui J, Gong K, Han P, et al (2020) Super Resolution of Arterial Spin Labeling MR Imaging Using Unsupervised Multi-Scale Generative Adversarial Network
- Pinto MF, Oliveira H, Batista S, Cruz L, Pinto M, et al. Prediction of disease progression and outcomes in multiple sclerosis with machine learning. Sci. rep. 2020;10:1-3.
- Ginat DT. Analysis of head CT scans flagged by deep learning software for acute intracranial hemorrhage. Neuroradiology. 2020;62:335-40.
- Kuo W, Hӓne C, Mukherjee P, Malik J, Yuh EL. Expert-level detection of acute intracranial hemorrhage on head computed tomography using deep learning. Proc. Natl. Acad. Sci. 2019;116:22737-45.
- Chang P, Grinband J, Weinberg BD, Bardis M, Khy M, et al. Deep-learning convolutional neural networks accurately classify genetic mutations in gliomas. Am J Neuroradiol. 2018; 39:1201-7.
- Cho J, Park KS, Karki M, Lee E, Ko S, et al. Improving sensitivity on identification and delineation of intracranial hemorrhage lesion using cascaded deep learning models. J. digit. Imaging. 2019; 32:450-61.
- Lee H, Yune S, Mansouri M, Kim M, Tajmir SH, et al. An explainable deep-learning algorithm for the detection of acute intracranial haemorrhage from small datasets. Nat. biomed. eng. 2019; 3:173-82.
- Ye H, Gao F, Yin Y, Guo D, Zhao P, et al. Precise diagnosis of intracranial hemorrhage and subtypes using a three-dimensional joint convolutional and recurrent neural network. Eur. radiol. 2019; 29:6191-201.
- Chilamkurthy S, Ghosh R, Tanamala S, Biviji M, Campeau NG, et al. Deep learning algorithms for detection of critical findings in head CT scans: a retrospective study. Lancet. 2018;392:2388-96.
- Yu Y, Xie Y, Thamm T, Gong E, Ouyang J, et al. Use of deep learning to predict final ischemic stroke lesions from initial magnetic resonance imaging. JAMA network open. 2020;3:e200772-.
- Ho KC, Scalzo F, Sarma KV, Speier W, El-Saden S, et al. Predicting ischemic stroke tissue fate using a deep convolutional neural network on source magnetic resonance perfusion images. J. Med Imaging. 2019;6:026001.
- J H, Jg Y, H P, et al (2019) Machine Learning-Based Model for Prediction of Outcomes in Acute Stroke. Stroke 50:. https://doi.org/10.1161/STROKEAHA.118.024293 [Google Scholar] [Crossref]
- Nielsen A, Hansen MB, Tietze A, Mouridsen K. Prediction of tissue outcome and assessment of treatment effect in acute ischemic stroke using deep learning. Stroke. 2018;49:1394-401.
- Murray NM, Unberath M, Hager GD, Hui FK. Artificial intelligence to diagnose ischemic stroke and identify large vessel occlusions: a systematic review. J. neurointerventional surg. 2020;12:156-64.
- Alawieh A, Zaraket F, Alawieh MB, Chatterjee AR, Spiotta A. Using machine learning to optimize selection of elderly patients for endovascular thrombectomy. J. NeuroInterventional Surg. 2019;11:847-51.
- Sichtermann T, Faron A, Sijben R, Teichert N, Freiherr J, et al. Deep learning–based detection of intracranial aneurysms in 3D TOF-MRA. Am. J. Neuroradiol. 2019;40:25-32.
- Park A, Chute C, Rajpurkar P, Lou J, Ball RL, et al. Deep learning–assisted diagnosis of cerebral aneurysms using the HeadXNet model. JAMA netw. open. 2019;2:e195600-.
- Stember JN, Chang P, Stember DM, Liu M, Grinband J, et al. Convolutional neural networks for the detection and measurement of cerebral aneurysms on magnetic resonance angiography. J digit imaging, 2019;32:808-15.
- Stone JR, Wilde EA, Taylor BA, Tate DF, Levin H, et al. Supervised learning technique for the automated identification of white matter hyperintensities in traumatic brain injury. Brain Injury. 2016; 30:1458-68.
- Jain S, Vyvere TV, Terzopoulos V, Sima DM, Roura E, et al. Automatic quantification of computed tomography features in acute traumatic brain injury. J. Neurotrauma. 2019;36:1794-803.
- Kerley CI, Huo Y, Chaganti S, Bao S, Patel MB, et al. Montage based 3D medical image retrieval from traumatic brain injury cohort using deep convolutional neural network. InMed. Imaging 2019: Image Process. 2019;10949:722-728.
- Katako A, Shelton P, Goertzen AL, et al (2018) Machine learning identified an Alzheimer’s disease-related FDG-PET pattern which is also expressed in Lewy body dementia and Parkinson’s disease dementia. Sci Rep 8:13236. https://doi.org/10.1038/s41598-018-31653-6 [Google Scholar] [Crossref]
- Fallahi A, Pooyan M, Lotfi N, Baniasad F, Tapak L, et al. Dynamic functional connectivity in temporal lobe epilepsy: a graph theoretical and machine learning approach. Neurological Sciences. 2021;42:2379-90.
- Choi KS, Choi SH, Jeong B. Prediction of IDH genotype in gliomas with dynamic susceptibility contrast perfusion MR imaging using an explainable recurrent neural network. Neuro-oncology. 2019;21:1197-209.
- Gaonkar B, Beckett J, Villaroman D, Ahn C, Edwards M, et al. Quantitative analysis of neural foramina in the lumbar spine: an imaging informatics and machine learning study. Radiol., Artif. Intell. 2019;1.
- Chang PD, Kuoy E, Grinband J, Weinberg BD, Thompson M, et al. Hybrid 3D/2D convolutional neural network for hemorrhage evaluation on head CT. Am. J. Neuroradiol. 2018;39:1609-16.
- Gao XW, Hui R, Tian Z. Classification of CT brain images based on deep learning networks. Comput. methods programs biomed. 2017;138:49-56.
- Grewal M, Srivastava MM, Kumar P, Varadarajan S. Radnet: Radiologist level accuracy using deep learning for hemorrhage detection in ct scans. In2018 IEEE 15th International Symposium on Biomedical Imaging 2018:281-284.
- Jnawali K, Arbabshirani MR, Rao N, Patel AA. Deep 3D convolution neural network for CT brain hemorrhage classification. InMed Imaging 2018: Computer-Aided Diagnosis 2018;10575: 307-313.
- Rudie JD, Rauschecker AM, Bryan RN, Davatzikos C, Mohan S. Emerging applications of artificial intelligence in neuro-oncology. Radiology. 2019;290:607.
- Ker IT. 3-Dimensional Convolutional Neural Network Diagnosis of Different Acute Brain Hemorrhages on Computed Tomography Scans. Sensors.19:2167.
- Flanders AE, Prevedello LM, Shih G, Halabi SS, Kalpathy-Cramer J, et al. RSNA-ASNR 2019 Brain Hemorrhage CT Annotators, Construction of a machine learning dataset through collaboration: the RSNA 2019 brain CT hemorrhage challenge, Radiology. Artif. Intell. 2020;2.
- Vlak MH, Algra A, Brandenburg R, Rinkel GJ. Prevalence of unruptured intracranial aneurysms, with emphasis on sex, age, comorbidity, country, and time period: a systematic review and meta-analysis. Lancet Neurol. 2011; 10:626-36.
- Wang L, Xu Q, Leung S, Chung J, Chen B, et al. Accurate automated Cobb angles estimation using multi-view extrapolation net. Med. Image Anal. 2019; 58:101542.
- Faron A, Sichtermann T, Teichert N, et al (2020) Performance of a Deep-Learning Neural Network to Detect Intracranial Aneurysms from 3D TOF-MRA Compared to Human Readers. Clin Neuroradiol 30:591–598. https://doi.org/10.1007/s00062-019-00809-w [Google Scholar] [Cross ref]
- Nakao T, Hanaoka S, Nomura Y, Sato I, Nemoto M, et al. Deep neural network‐based computer‐assisted detection of cerebral aneurysms in MR angiography. J. Magn. Reson. Imaging. 2018;47:948-53.
- Yang X, Xia D, Kin T, Igarashi T. Intra: 3d intracranial aneurysm dataset for deep learning. InProceedings IEEE/CVF Conf. Comput. Vis. Pattern Recognit. 2020:2656-2666.
- You J, Yu PL, Tsang AC, Tsui EL, Woo PP, et al. Automated computer evaluation of acute ischemic stroke and large vessel occlusion. arXiv preprint arXiv:1906.08059. 2019.
- Takahashi N, Lee Y, Tsai DY, Matsuyama E, Kinoshita T, et al. An automated detection method for the MCA dot sign of acute stroke in unenhanced CT. Radiological physics and technology. 2014; 7:79-88.
- Chen Z, Zhang R, Xu F, Gong X, Shi F, et al. Novel prehospital prediction model of large vessel occlusion using artificial neural network. Front aging neurosci. 2016;10:181.
- Abstract WMP16: Artificial Intelligence Detection of Cerebrovascular Large Vessel Occlusion - Nine Month, 650 Patient Evaluation of the Diagnostic Accuracy and Performance of the Viz.ai LVO Algorithm Arindam Chatterjee, Nayana R Somayaji, Ismail M Kabakis
- Chen L, Bentley P, Rueckert D. Fully automatic acute ischemic lesion segmentation in DWI using convolutional neural networks. NeuroImage: Clin. 2017;15:633-43.
- Guerrero R, Qin C, Oktay O, Bowles C, Chen L, et al. White matter hyperintensity and stroke lesion segmentation and differentiation using convolutional neural networks. NeuroImage: Clin. 2018;17:918-34.
- Öman O, Mäkelä T, Salli E, Savolainen S, Kangasniemi M. 3D convolutional neural networks applied to CT angiography in the detection of acute ischemic stroke. Eur radiol exp. 2019;3:1-1.
- Qiu W, Kuang H, Teleg E, Ospel JM, Sohn SI, Et al. Machine learning for detecting early infarction in acute stroke with non–contrast-enhanced CT. Radiology. 2020;294:638-44.
- Barber PA, Demchuk AM, Zhang J, Buchan AM. Validity and reliability of a quantitative computed tomography score in predicting outcome of hyperacute stroke before thrombolytic therapy. The Lancet. 2000 ;355(9216):1670-4.
- Goebel J, Stenzel E, Guberina N, Wanke I, Koehrmann M, et al. Automated ASPECT rating: comparison between the Frontier ASPECT Score software and the Brainomix software. Neuroradiology.2018;60:1267-72 .
- Guberina N, Dietrich U, Radbruch A, Goebel J, Deuschl C,et al. Detection of early infarction signs with machine learning-based diagnosis by means of the Alberta Stroke Program Early CT score (ASPECTS) in the clinical routine. Neuroradiology.2018;60:889-901.
- Maegerlein C, Fischer J, Mönch S, Berndt M, Wunderlich S,et al. Automated calculation of the Alberta Stroke Program Early CT Score: feasibility and reliability. Radiology.2019;291:141-148.
- Seker F, Pfaff J, Nagel S, Vollherbst D, Gerry S,et al. CT reconstruction levels affect automated and reader‐based ASPECTS ratings in acute ischemic stroke. J Neuroimaging.2019;29:62-64.
- Nishi H, Oishi N, Ishii A, Ono I, Ogura T,et al.Deep learning–derived high-level neuroimaging features predict clinical outcomes for large vessel occlusion.Stroke.2020 ;51:1484-92.
- Nair T, Precup D, Arnold DL, Arbel T. Exploring uncertainty measures in deep networks for multiple sclerosis lesion detection and segmentation. Med image anal. 2020;59:101557.
- Wang SH, Tang C, Sun J, Yang J, Huang C,et al. Multiple sclerosis identification by 14-layer convolutional neural network with batch normalization, dropout, and stochastic pooling. Front neurosci.2018;12:1-11.
- Yoo Y, Tang LY, Li DK, Metz L, Kolind S,et al. Deep learning of brain lesion patterns and user-defined clinical and MRI features for predicting conversion to multiple sclerosis from clinically isolated syndrome. Comput Methods Biomech Biomed Eng Imaging Vis. 2019;7:250-259.
- Guo BJ, Yang ZL, Zhang LJ. Gadolinium deposition in brain: current scientific evidence and future perspectives. Frontiers in molecular neuroscience. 2018;11:335.
- Narayana PA, Coronado I, Sujit SJ, Wolinsky JS, Lublin FD,et al . Deep learning for predicting enhancing lesions in multiple sclerosis from noncontrast MRI. Radiology.2020;294:398-404.
- Tomita N, Cheung YY, Hassanpour S. Deep neural networks for automatic detection of osteoporotic vertebral fractures on CT scans. Comput biol Med 2018;98:8-15.
- Bar A, Wolf L, Amitai OB, Toledano E, Elnekave E. Compression fractures detection on CT.In Medical imaging.2017;10134:1036-1043
- Derkatch S, Kirby C, Kimelman D, Jozani MJ, Davidson JM,et al. Identification of vertebral fractures by convolutional neural networks to predict nonvertebral and hip fractures: a registry-based cohort study of dual X-ray absorptiometry.Radiology.2019;293:405-411.
- Baid U, Shah NA, Talbar S. Brain tumor segmentation with cascaded deep convolutional neural network.2020 90-98. Springer Int Publ
- Zhou C, Ding C, Wang X, Lu Z, Tao D. One-pass multi-task networks with cross-task guided attention for brain tumor segmentation. IEEE Trans Image Process 2020;29:4516-4529.
- Mlynarski P, Delingette H, Criminisi A, Ayache N. Deep learning with mixed supervision for brain tumor segmentation. J Med Imaging. 2019;6:034002.
- Chen S, Ding C, Liu M. Dual-force convolutional neural networks for accurate brain tumor segmentation. Pattern Recognition. 2019 ;88:90-100.
- Pereira S, Pinto A, Alves V, Silva CA. Brain tumor segmentation using convolutional neural networks in MRI images. IEEE trans. med. imaging. 2016;35:1240-1251.
- Parsons D W, Jones S,Zhang X, Cheng-Ho Lin J, Leary R J, Angenendt P, Mankoo P, et al An integrated genomic analysis of human glioblastoma multiforme. Science.2008; 321:1807–1812.
- Zhao J, Huang Y, Song Y, Xie D, Hu M, et al.Diagnostic accuracy and potential covariates for machine learning to identify IDH mutations in glioma patients: evidence from a meta-analysis. Eur Radiol. 2020;30:4664-4674.
- Haubold J, Demircioglu A, Gratz M, Glas M, Wrede K,et al. Non-invasive tumor decoding and phenotyping of cerebral gliomas utilizing multiparametric 18F-FET PET-MRI and MR Fingerprinting. Eur J Nucl Med Mol. Imaging.2020;47:1435-1445.
- Li H, Habes M, Wolk DA, Fan Y. A deep learning model for early prediction of Alzheimer’s disease dementia based on hippocampal MRI.1904; 555 1-40.
- Basaia S, Agosta F, Wagner L, Canu E, Magnani G, et al. Alzheimer's Disease Neuroimaging Initiative. Automated classification of Alzheimer's disease and mild cognitive impairment using a single MRI and deep neural networks. NeuroImage: Clinical. 2019; 21:101645.
- Young AL, Marinescu RV, Oxtoby NP, Bocchetta M, Yong K,et al. Uncovering the heterogeneity and temporal complexity of neurodegenerative diseases with Subtype and Stage Inference. Nat. commun. 2018;9:4273. [Google scholar]
- Spasov S, Passamonti L, Duggento A, Liò P, Toschi N. A Parameter-Efficient Deep Learning Approach To Predict Conversion From Mild Cognitive Impairment To Alzheimer’s Disease. Neuroimage. 2019;189:276-287.[Google scholar]
- Ashton NJ, Nevado-Holgado AJ, Barber IS, Lynham S, Gupta V,et al. A plasma protein classifier for predicting amyloid burden for preclinical Alzheimer’s disease. Sci. adv. 2019 ;5:7220.
- Benjamin Goudey , Bowen J Fung , Christine Schieber , Noel G Faux. A blood-based signature of cerebrospinal fluid Aβ1-42 status.Sci Rep 2019; 9:4163–4163.
- Wang L, Sha L, Lakin JR, Bynum J, Bates DW, et al Development and validation of a deep learning algorithm for mortality prediction in selecting patients with dementia for earlier palliative care interventions. JAMA netw. open. 2019;2:1-14
- Tang Z, Chuang KV, DeCarli C, Jin LW, Beckett L,et al. Interpretable classification of Alzheimer’s disease pathologies with a convolutional neural network pipeline. Nat. commun. 2019;10:1-4.
- De Tommaso M, De Carlo F, Difruscolo O, Massafra R, Sciruicchio V,et al. Detection of subclinical brain electrical activity changes in Huntington's disease using artificial neural networks. Clin. neurophysio. 2003 ;114:1237-1245.
- Odish OF, Johnsen K, van Someren P, Roos RA, van Dijk JG. EEG may serve as a biomarker in Huntington’s disease using machine learning automatic classification. Sci rep. 2018;8:1-8.
- De Tommaso M, De Carlo F, Difruscolo O, Massafra R, Sciruicchio V,et al. Detection of subclinical brain electrical activity changes in Huntington's disease using artificial neural networks. Clin neurophysiol. 2003 ;114:1237-45.[Google scholar]
- Benbadis SR, Beniczky S, Bertram E, MacIver S, Moshé SL. The role of EEG in patients with suspected epilepsy. Epileptic Disorders. 2020;22:143-155.[Google scholar]
- Park YW, Choi YS, Kim SE, Choi D, Han K,et al. Radiomics features of hippocampal regions in magnetic resonance imaging can differentiate medial temporal lobe epilepsy patients from healthy controls. Sci rep. 2020; 10:1-8.
- Mo J, Liu Z, Sun K, Ma Y, Hu W, et al.Automated detection of hippocampal sclerosis using clinically empirical and radiomics features. Epilepsia.2019; 60:2519-2529.
- Beheshti I, Sone D, Maikusa N, Kimura Y, Shigemoto Y,et al. Pattern analysis of glucose metabolic brain data for lateralization of MRI-negative temporal lobe epilepsy. Epilepsy Res. 2020;167:106474.
- Sahebzamani G, Saffar M, Soltanian-Zadeh H. Machine learning based analysis of structural MRI for epilepsy diagnosis. In2019 4th Int. Conf. Pattern Recognit. Image Anal. (IPRIA) 2019; 58-63.
- Mahmoudi F, Elisevich K, Bagher-Ebadian H, Nazem-Zadeh MR,et al.Data mining MR image features of select structures for lateralization of mesial temporal lobe epilepsy. Plos one. 2018;13:e0199137.
- Gleichgerrcht E, Munsell BC, Alhusaini S, Alvim MK, Bargalló N,et al. Artificial intelligence for classification of temporal lobe epilepsy with ROI-level MRI data: A worldwide ENIGMA-Epilepsy study. NeuroImage: Clin. 2021;31:102765.
- Kabat J, Król P. Focal cortical dysplasia–review. Pol. j. radiol. 2012; 77:35.
- Wang H, Ahmed SN, Mandal M. Automated detection of focal cortical dysplasia using a deep convolutional neural network. Comput. Med. Imaging Graph. 2020;79:101662.
- Jin B, Krishnan B, Adler S, Wagstyl K, Hu W,et al. Automated detection of focal cortical dysplasia type II with surface‐based magnetic resonance imaging postprocessing and machine learning. Epilepsia.2018 ;59:982-92.
- Hwang G, Hermann B, Nair VA, Conant LL, Dabbs K,et al. Brain aging in temporal lobe epilepsy: Chronological, structural, and functional.NeuroImage: Clinical. 2020 ;25:102183.
- La Rocca M, Garner R, Jann K, Kim H, Vespa P,et al. Machine learning of multimodal MRI to predict the development of epileptic seizures after traumatic brain injury. In Int. Conf. Med. Imaging Deep Learn.2019.
- Roy T, Pandit A. Neuroimaging in epilepsy. Ann Indian Acad Neurol 2011;14:14:78.
- Mazrooyisebdani M, Nair VA, Garcia-Ramos C, Mohanty R, Meyerand E, et al. Graph theory analysis of functional connectivity combined with machine learning approaches demonstrates widespread network differences and predicts clinical variables in temporal lobe epilepsy. Brain connectivity. 2020;10:39-50.
- Hekmati R, Azencott R, Zhang W, Chu ZD, Paldino MJ. Localization of epileptic seizure focus by computerized analysis of fMRI recordings. Brain Inform. 2020; 7:1-3.
- Hwang G, Nair VA, Mathis J, Cook CJ, Mohanty R,et al. Using low-frequency oscillations to detect temporal lobe epilepsy with machine learning. Brain connectivity. 2019;9:184-93.
- Bharath RD, Panda R, Raj J, Bhardwaj S, Sinha S,et al. Machine learning identifies “rsfMRI epilepsy networks” in temporal lobe epilepsy. Eur. radiol.2019; 29:3496-3505.
- Szaflarski JP, Gloss D, Binder JR, Gaillard WD, Golby AJ,et al.Practice guideline summary: Use of fMRI in the presurgical evaluation of patients with epilepsy: Report of the Guideline Development, Dissemination, and Implementation Subcommittee of the American Academy of Neurology. Neurology. 2017; 88:395-402.
- Bauer PR, Reitsma JB, Houweling BM, Ferrier CH, Ramsey NF. Can fMRI safely replace the Wada test for preoperative assessment of language lateralisation? A meta-analysis and systematic review. J. Neurol. Neurosurg. Psychiatry. 2014;85:581-588.
- Sollee J, Tang L, Igiraneza AB, Xiao B, Bai HX,et al. Artificial intelligence for medical image analysis in epilepsy. Epilepsy Res. 2022:106861.
- Jamaludin A, Lootus M , Kadir T , Zisserman A , Urban, J,et al, ISSLS PRIZE IN BIOENGINEERING SCIENCE 2017: Automation of reading of radiological features from magnetic resonance images (MRIs) of the lumbar spine without human intervention is comparable with an expert radiologist. Eur. spine j. 2017;26,1374-1383.
- Chen B, Xu Q, Wang L, Leung S, Chung J, Li S. An automated and accurate spine curve analysis system. Ieee Access. 2019;7:124596-124605.
- Huang J, Shen H, Wu J, Hu X, Zhu Z,et al. Spine Explorer: a deep learning based fully automated program for efficient and reliable quantifications of the vertebrae and discs on sagittal lumbar spine MR images. Spine J. 2020;20:590-599.
- Whitehead W, Moran S, Gaonkar B, Macyszyn L, Iyer S. A deep learning approach to spine segmentation using a feed-forward chain of pixel-wise convolutional networks. IEEE 15th Int. Symp Biomed Imaging 2018; 868-871.
- Kim BS, Yu M, Kim S, Yoon JS, Baek S. Scale-attentional U-Net for the segmentation of the median nerve in ultrasound images. Ultrasonography. 2022; 41:706-717.
- Hareendranathan AR, Chahal BS, Zonoobi D, Sukhdeep D, Jaremko JL. Artificial intelligence to automatically assess scan quality in hip ultrasound. Indian J Orthop 2020;55:1535-1542.
- Lloyd J, Morse R, Taylor A, Phillips D, Higham H,et al. Artificial Intelligence: Innovation to Assist in the Identification of Sono-anatomy for Ultrasound-Guided Regional Anaesthesia. Biomed Vis 2022:117-40.
- McKendrick M, Yang S, McLeod GA. The use of artificial intelligence and robotics in regional anaesthesia. Anaesthesia. 2021;76:171-81.
- Suk HI, Liu M, Yan P, Lian C. Machine Learning in Medical Imaging: 10th International Workshop, MLMI 2019, Held in Conjunction with MICCAI 2019. Springer Nature; 2019.
- Bowness J, Varsou O, Turbitt L, Burkett‐St Laurent D. Identifying anatomical structures on ultrasound: assistive artificial intelligence in ultrasound‐guided regional anesthesia. Clin. Anat. 2021;34:802-809.
- Nguyen G K, Shetty A, Artificial Intelligence and Machine Learning: Opportunities for Radiologists in Training J Am Coll Radiol. 2018;15:1320-1321.
- Duong MT, Rauschecker AM, Rudie JD, Chen PH, Cook TS,et al. Artificial intelligence for precision education in radiology. Br j radiol. 2019;92:20190389.
- Bowness J, El‐Boghdadly K, Burckett‐St Laurent D. Artificial intelligence for image interpretation in ultrasound-guided regional anaesthesia. Anaesthesia. 2021;76:602-607.